Extensive, multifactorial data sharing is a crucial prerequisite for current and future (radiotherapy) research. However, the cost, time and effort to achieve this are often a roadblock. We present an open-source based data-sharing infrastructure between two radiotherapy departments, allowing seamless exchange of de-identified, automatically translated clinical and biomedical treatment data.
Legal and ethical
A collaboration and data transfer agreement was signed which describes the type of data, the permitted use and the protection of the data. This agreement was submitted to and approved by the local ethical authority.
An example agreement is attached below.
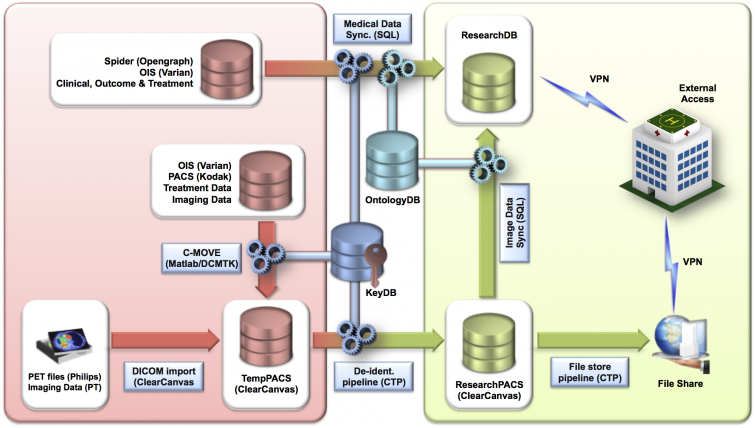
Overview of data sources, flow and external access. Terms as mentioned in the table below:
| Term | Meaning |
|---|---|
| CTP | Clinical Trial Processor |
| KeyDB | Secure key database linking random research patient identifier (ResearchID) and original patient data |
| OntologyGB | Database with preferred terms and concept IDs after mapping local terms to SNOMED CT concepts. |
| ResearchDB | Research database holding medical data and imaging meta data |
| ResearchID | Random patient identification code |
| ResearchPACS | Research PACS partition holding only de-identified DICOM objects |
| SeriesInstanceUID | Unique series identifier for all images in a series for a given patient` |
| SNOMED-CT | Systematized Nomenclature of Medicine - Clinical Terms |
| TempPACS | Temporary PACS partition holding identifiable DICOM headers |
| UID | Unique IDentifier |
| Attachment | Size |
|---|---|
| 40.5 KB |